Register for free to join our community of investors and share your ideas. You will also get access to streaming quotes, interactive charts, trades, portfolio, live options flow and more tools.
1 left 0007 hitting
12 milion bid vs. 2 milion ask BANGGGGG COMING EOD
big eod buying pressure for sure. thats not all today. thats for sure. mm bid 10 milion waiting for bid dumper. but noone sell. and before close he must load more out of ask. i am sure. DNAG will blowing...
something big arround the corner. watch the bid. wow
i hold few months here. and now banggg. i think this just the begining here
looks great here. suddenly big volume. someone load here. next week will be great
I just realized that all those post came from one person. Got in at 5's whish I could have been here at 2's
Here with you brother, this will be in .001 soon
tic toc... 0007 will falling. no one sell it. looks like someone load up. why ever reason. DNAG going to fly
just 1 0007 hit and 0008 up blue skies... crazy. whata going on here.
hey i am alone in here? big eod for sure. 001 break coming
25 milion trading volume so far. strong... 0007 falling and then bangggggg
last 1.5 milion and 0008 then 0011 up
DNAG MAKING BANGGGGGGGGGGGGGGGGGGGGGGGG 002 EOD
250%+ and going. blow up this afternoon. next week 005-01 with big news
10 milion bid order coming in CRAZYYYYYYYYYYYYYYYYYYYYYYYYYY
DNAG BLOWING UP............. 0007 1 left
200%+ and no one in here? will blow up
just 1 milion left to 001 break crazy
monster volume coming in
0006 up 1 left and huge bid 0005 builing. wowwwwwwwwwwwwwwwww
looks like 001 break coming quick
0005 1 left big volume coming in.
0006 coming. whats up here?
crazy bids 0004
big hits coming in. 0004 falling now
Agree, looking good here. Every day smart 0003 hits. Looks like someone collect smart. Still waiting for huge breakout soon. Stay long
Thanks Hunter. I'm still hanging with DNAG. I think something is coming up soon. I'm going to add a million shares, they're still cheap enough.
could be. i collect some bottom shares and i am long here. with this low share structure big potential here. R/M - any good news and easy 10-30 bagger from here. buy cheap hold long. that it is. keep an eye on it. more and more smart ask hits last few weeks.
What do you think ? Will this come back from the dead
awesome 0003 trading volume yesterday.
0003 up nice volume early morning
Havent seen UBSS on the lead bid! Nite took over with 5 million after ubss got 1.4 partial.
|
Followers
|
297
|
Posters
|
|
|
Posts (Today)
|
0
|
Posts (Total)
|
82595
|
|
Created
|
08/19/00
|
Type
|
Free
|
| Moderators | |||
DNA
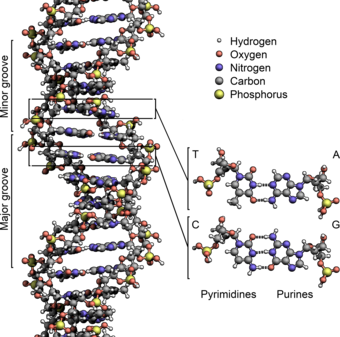

Deoxyribonucleic acid (DNA) is a molecule encoding the genetic instructions used in the development and functioning of all known living organisms and many viruses. Along with RNA and proteins, DNA is one of the three major macromolecules that are essential for all known forms of life. Genetic information is encoded as a sequence of nucleotides (guanine, adenine, thymine, and cytosine) recorded using the letters G, A, T, and C. Most DNA molecules are double-stranded helices, consisting of two long polymers of simple units called nucleotides, molecules with backbones made of alternating sugars (deoxyribose) and phosphate groups (related to phosphoric acid), with the nucleobases (G, A, T, C) attached to the sugars. DNA is well-suited for biological information storage, since the DNA backbone is resistant to cleavage and the double-stranded structure provides the molecule with a built-in duplicate of the encoded information.
These two strands run in opposite directions to each other and are therefore anti-parallel, one backbone being 3' (three prime) and the other 5' (five prime). This refers to the direction the 3rd and 5th carbon on the sugar molecule is facing. Attached to each sugar is one of four types of molecules called nucleobases (informally, bases). It is the sequence of these four nucleobases along the backbone that encodes information. This information is read using the genetic code, which specifies the sequence of the amino acids within proteins. The code is read by copying stretches of DNA into the related nucleic acid RNA in a process called transcription.
Within cells, DNA is organized into long structures called chromosomes. During cell division these chromosomes are duplicated in the process of DNA replication, providing each cell its own complete set of chromosomes. Eukaryotic organisms (animals, plants, fungi, and protists) store most of their DNA inside the cell nucleus and some of their DNA in organelles, such as mitochondria or chloroplasts.[1] In contrast, prokaryotes (bacteria and archaea) store their DNA only in the cytoplasm. Within the chromosomes, chromatin proteins such as histones compact and organize DNA. These compact structures guide the interactions between DNA and other proteins, helping control which parts of the DNA are transcribed.
Properties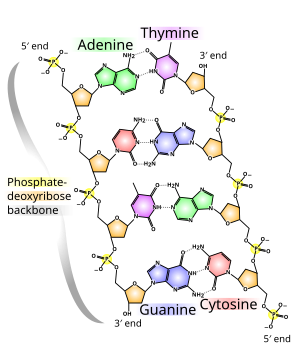
DNA is a long polymer made from repeating units called nucleotides.[2][3][4] DNA was first identified and isolated by Friedrich Miescher and the double helix structure of DNA was first discovered by James D. Watson and Francis Crick. The structure of DNA of all species comprises two helical chains each coiled round the same axis, and each with a pitch of 34 ångströms (3.4 nanometres) and a radius of 10 ångströms (1.0 nanometres).[5] According to another study, when measured in a particular solution, the DNA chain measured 22 to 26 ångströms wide (2.2 to 2.6 nanometres), and one nucleotide unit measured 3.3 Å (0.33 nm) long.[6] Although each individual repeating unit is very small, DNA polymers can be very large molecules containing millions of nucleotides. For instance, the largest human chromosome, chromosome number 1, is approximately 220 million base pairs long.[7]
In living organisms DNA does not usually exist as a single molecule, but instead as a pair of molecules that are held tightly together.[8][9] These two long strands entwine like vines, in the shape of a double helix. The nucleotide repeats contain both the segment of the backbone of the molecule, which holds the chain together, and a nucleobase, which interacts with the other DNA strand in the helix. A nucleobase linked to a sugar is called a nucleoside and a base linked to a sugar and one or more phosphate groups is called a nucleotide. A polymer comprising multiple linked nucleotides (as in DNA) is called a polynucleotide.[10]
The backbone of the DNA strand is made from alternating phosphate and sugar residues.[11] The sugar in DNA is 2-deoxyribose, which is a pentose (five-carbon) sugar. The sugars are joined together by phosphate groups that form phosphodiester bonds between the third and fifth carbon atoms of adjacent sugar rings. These asymmetric bonds mean a strand of DNA has a direction. In a double helix the direction of the nucleotides in one strand is opposite to their direction in the other strand: the strands are antiparallel. The asymmetric ends of DNA strands are called the 5′ (five prime) and 3′ (three prime) ends, with the 5' end having a terminal phosphate group and the 3' end a terminal hydroxyl group. One major difference between DNA and RNA is the sugar, with the 2-deoxyribose in DNA being replaced by the alternative pentose sugar ribose in RNA.[9]
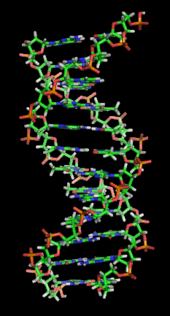
The DNA double helix is stabilized primarily by two forces: hydrogen bonds between nucleotides and base-stacking interactions among aromatic nucleobases.[13] In the aqueous environment of the cell, the conjugated π bonds of nucleotide bases align perpendicular to the axis of the DNA molecule, minimizing their interaction with the solvation shell and therefore, the Gibbs free energy. The four bases found in DNA are adenine (abbreviated A), cytosine (C), guanine (G) and thymine (T). These four bases are attached to the sugar/phosphate to form the complete nucleotide, as shown for adenosine monophosphate.
The nucleobases are classified into two types: the purines, A and G, being fused five- and six-membered heterocyclic compounds, and the pyrimidines, the six-membered rings C and T.[9] A fifth pyrimidine nucleobase, uracil (U), usually takes the place of thymine in RNA and differs from thymine by lacking a methyl group on its ring. In addition to RNA and DNA a large number of artificial nucleic acid analogues have also been created to study the properties of nucleic acids, or for use in biotechnology.[14]
Uracil is not usually found in DNA, occurring only as a breakdown product of cytosine. However in a number of bacteriophages - Bacillus subtilis bacteriophages PBS1 and PBS2 and Yersinia bacteriophage piR1-37 - thymine has been replaced by uracil.[15] A modified form (beta-d-glucopyranosyloxymethyluracil) is also found in a number of organisms: the flagellates Diplonema and Euglena, and all the kinetoplastid genera[16] Biosynthesis of J occurs in two steps: in the first step a specific thymidine in DNA is converted into hydroxymethyldeoxyuridine; in the second HOMedU is glycosylated to form J.[17] Proteins that bind specifically to this base have been identified.[18][19][20] These proteins appear to be distant relatives of the Tet1 oncogene that is involved in the pathogenesis of acute myeloid leukemia.[21] J appears to act as a termination signal for RNA polymerase II.[22][23]
| Volume | |
| Day Range: | |
| Bid Price | |
| Ask Price | |
| Last Trade Time: |
